

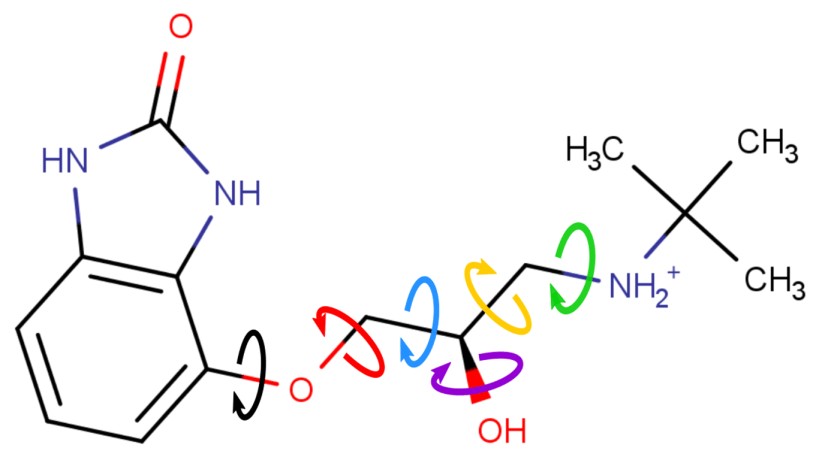
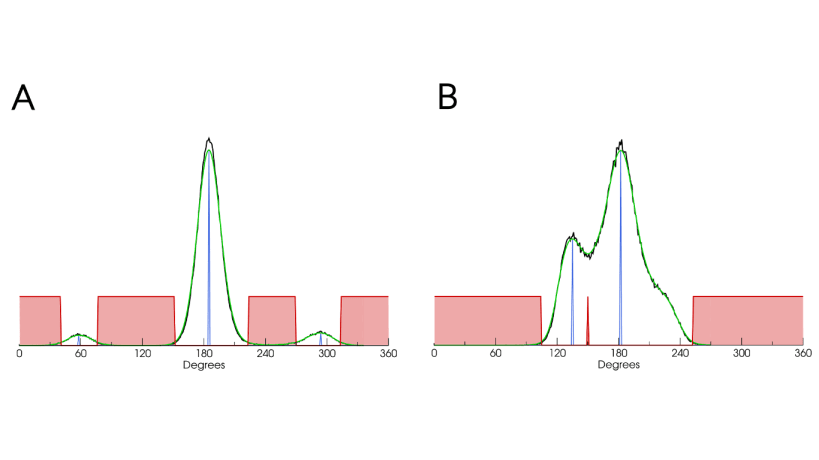
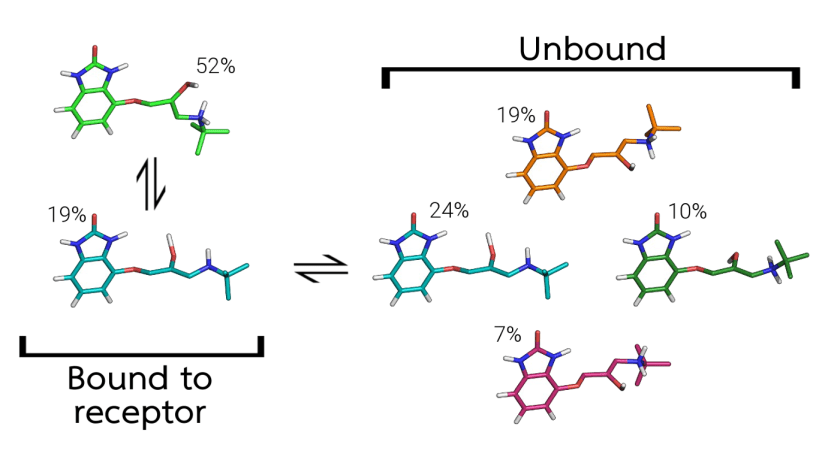
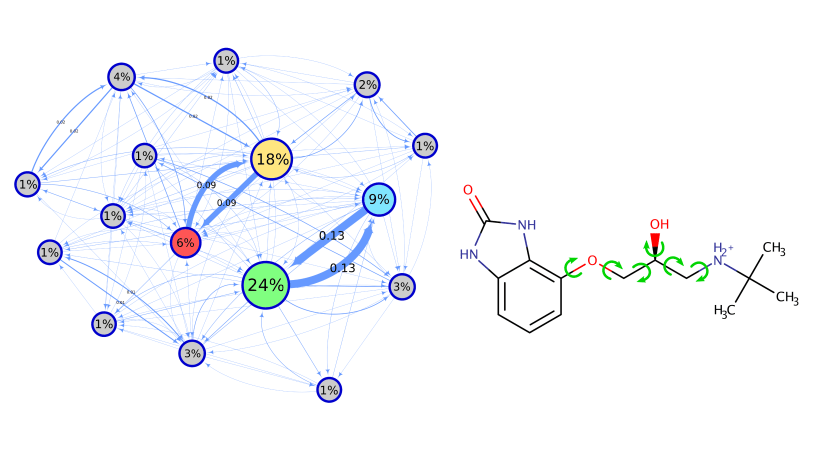
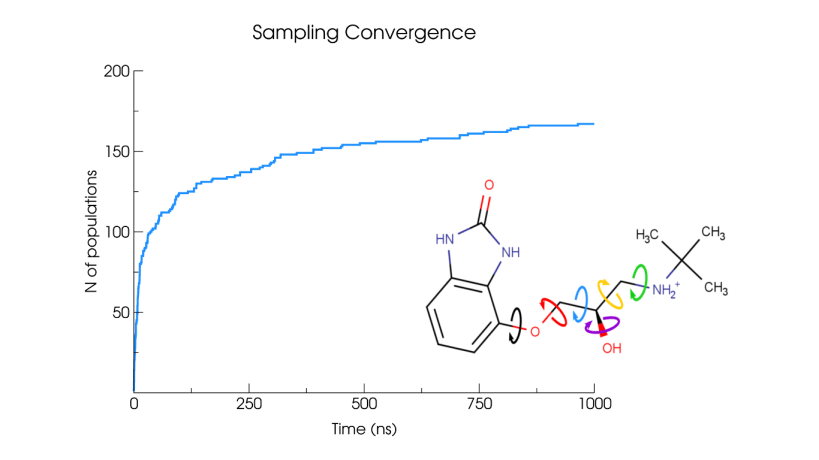
Conformational generation is a recurrent challenge in early phases of drug design, mostly due to the task of making sense between the number of conformers generated and their relevance for biological purposes. In this sense, ConfID, a Python-based computational tool, was designed to identify and characterize conformational populations of drug-like molecules sampled through molecular dynamics simulations. By using molecular dynamics (MD) simulations (and assuming accurate parameters are used), ConfID can identify all conformational populations sampled in the presence of solvent and quantify their relative abundance, while harnessing the benefits of MD and calculating time-dependent properties of each conformational population identified. To read a complete ConfID documentation and tutorials, access: https://github.com/sbcblab/confid
This snap hasn't been updated in a while. It might be unmaintained and have stability or security issues.
You are about to open
Do you wish to proceed?
Thank you for your report. Information you provided will help us investigate further.
There was an error while sending your report. Please try again later.
Snaps are applications packaged with all their dependencies to run on all popular Linux distributions from a single build. They update automatically and roll back gracefully.
Snaps are discoverable and installable from the Snap Store, an app store with an audience of millions.

Snap can be installed from the command line on openSUSE Leap 15.x and Tumbleweed.
You need first add the snappy repository from the terminal. Choose the appropriate command depending on your installed openSUSE flavor.
Tumbleweed:
sudo zypper addrepo --refresh https://download.opensuse.org/repositories/system:/snappy/openSUSE_Tumbleweed snappy
Leap 15.x:
sudo zypper addrepo --refresh https://download.opensuse.org/repositories/system:/snappy/openSUSE_Leap_15.6 snappy
If needed, Swap out openSUSE_Leap_15. for, openSUSE_Leap_16.0 if you’re using a different version of openSUSE.
With the repository added, import its GPG key:
sudo zypper --gpg-auto-import-keys refresh
Finally, upgrade the package cache to include the new snappy repository:
sudo zypper dup --from snappy
Snap can now be installed with the following:
sudo zypper install snapd
You then need to either reboot, logout/login or source /etc/profile to have /snap/bin added to PATH.
Additionally, enable and start both the snapd and the snapd.apparmor services with the following commands:
sudo systemctl enable --now snapd
sudo systemctl enable --now snapd.apparmor
To install ConfID: Conformational Identifier, simply use the following command:
sudo snap install confid
Browse and find snaps from the convenience of your desktop using the snap store snap.

Interested to find out more about snaps? Want to publish your own application? Visit snapcraft.io now.