

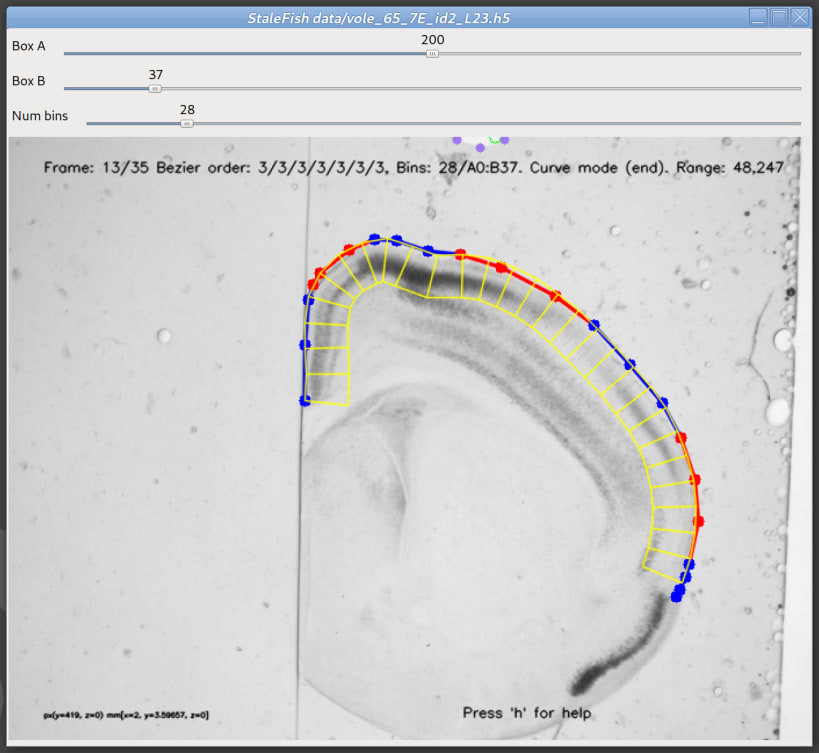
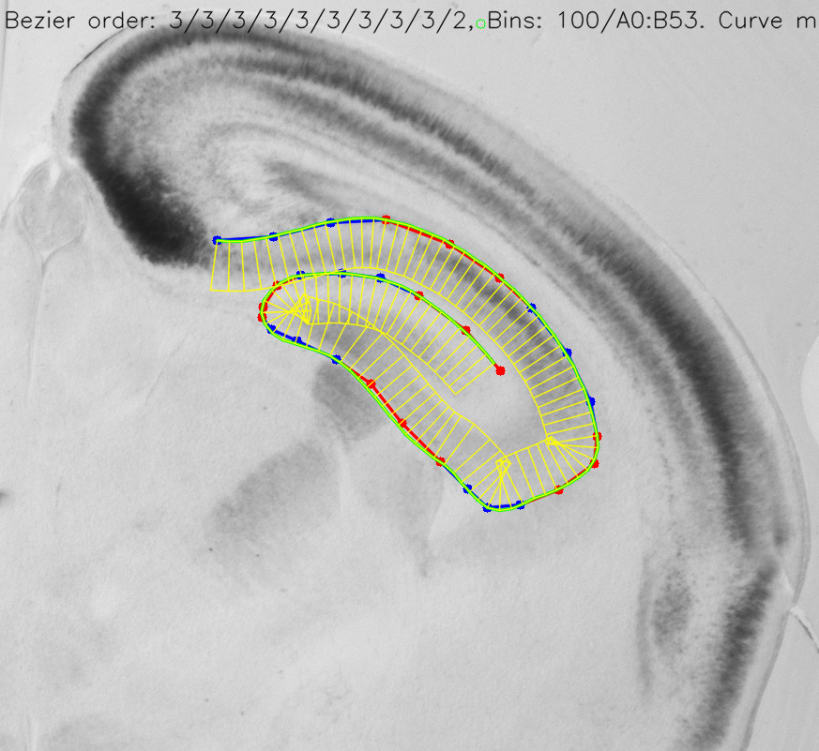
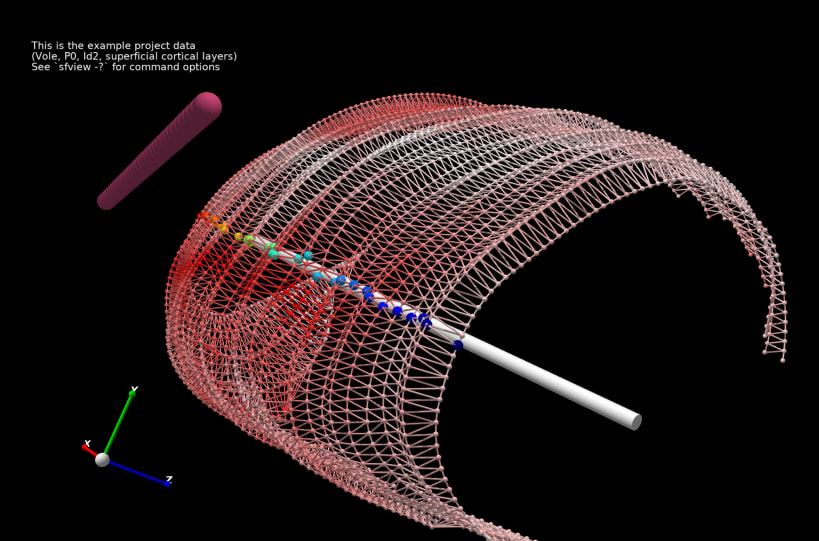
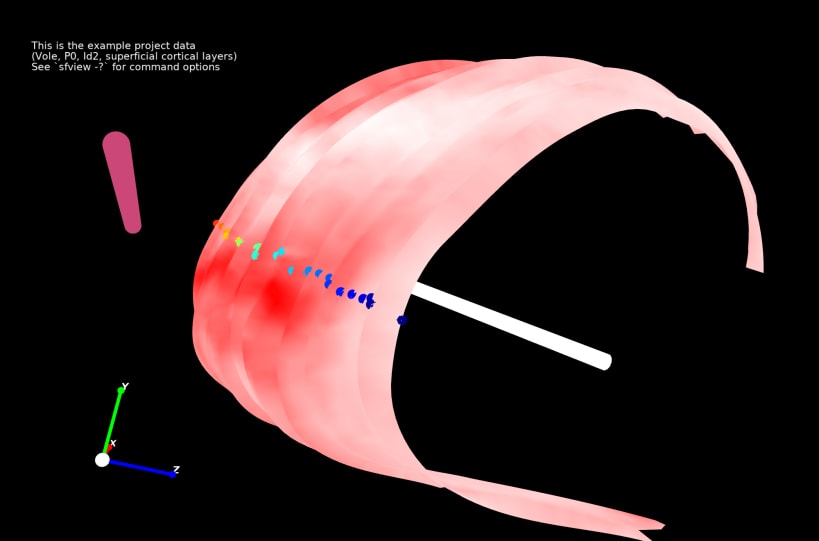
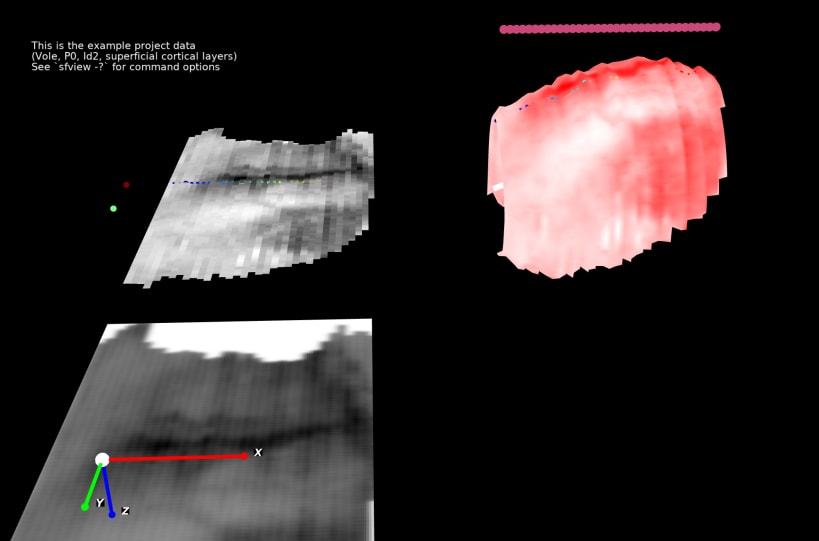
Stalefish is a tool to analyse gene expression. From your brain slice images you can build 3D expression surfaces and then convert these into 2D expression maps.
For more details see these slides: https://bit.ly/3At1yrv and the paper https://www.pnas.org/doi/10.1073/pnas.2113896119
You are about to open
Do you wish to proceed?
Thank you for your report. Information you provided will help us investigate further.
There was an error while sending your report. Please try again later.
Snaps are applications packaged with all their dependencies to run on all popular Linux distributions from a single build. They update automatically and roll back gracefully.
Snaps are discoverable and installable from the Snap Store, an app store with an audience of millions.

Snap is available for Red Hat Enterprise Linux (RHEL) 8 and RHEL 7, from the 7.6 release onward.
The packages for RHEL 7, RHEL 8, and RHEL 9 are in each distribution’s respective Extra Packages for Enterprise Linux (EPEL) repository. The instructions for adding this repository diverge slightly between RHEL 7, RHEL 8 and RHEL 9, which is why they’re listed separately below.
The EPEL repository can be added to RHEL 9 with the following command:
sudo dnf install https://dl.fedoraproject.org/pub/epel/epel-release-latest-9.noarch.rpm
sudo dnf upgrade
The EPEL repository can be added to RHEL 8 with the following command:
sudo dnf install https://dl.fedoraproject.org/pub/epel/epel-release-latest-8.noarch.rpm
sudo dnf upgrade
The EPEL repository can be added to RHEL 7 with the following command:
sudo rpm -ivh https://dl.fedoraproject.org/pub/epel/epel-release-latest-7.noarch.rpm
Adding the optional and extras repositories is also recommended:
sudo subscription-manager repos --enable "rhel-*-optional-rpms" --enable "rhel-*-extras-rpms"
sudo yum update
Snap can now be installed as follows:
sudo yum install snapd
Once installed, the systemd unit that manages the main snap communication socket needs to be enabled:
sudo systemctl enable --now snapd.socket
To enable classic snap support, enter the following to create a symbolic link between /var/lib/snapd/snap and /snap:
sudo ln -s /var/lib/snapd/snap /snap
Either log out and back in again or restart your system to ensure snap’s paths are updated correctly.
To install Stalefish, simply use the following command:
sudo snap install stalefish
Browse and find snaps from the convenience of your desktop using the snap store snap.

Interested to find out more about snaps? Want to publish your own application? Visit snapcraft.io now.